Kernel Regularized Least Squares — an Explainer
How KRLS works, when to use it, and how to run it in R and Stata.
This is a self-contained tutorial on Kernel Regularized Least Squares (KRLS) for users coming from either R or Stata. The R commands use the KRLS package; the Stata commands use the krls command. Both implement the same algorithm from Hainmueller and Hazlett (2014) and Ferwerda, Hainmueller, and Hazlett (2017).
← Back to the Kernel ML Methods project page for the package landing pages and the most recent release notes.
Do not use this when: the dataset is very large, pure prediction is the goal, or the estimand requires extrapolation far outside the observed covariate space.
The problem KRLS solves
You want to estimate the conditional expectation \(E[Y \mid X]\) without committing to a specific functional form. The standard options each have their own failure modes:
- OLS assumes additivity and linearity. If either is wrong, the coefficients are biased and the marginal effects (the things you actually care about) are wrong everywhere.
- GLM with interactions and polynomials can pick up non-linearities, but you have to specify them in advance. With \(p\) predictors, the cost of “just include the right interactions” grows fast.
- Random forests / gradient boosting are flexible but not interpretable: you get predictions, not coefficients, and pointwise marginal effects are not first-class output.
- GAMs assume additivity, which can be too restrictive for social-science data with rich interactions.
KRLS is a kernel-based method that:
- Makes no assumption about the functional form (no linearity, no additivity, no specific interactions).
- Returns pointwise marginal effects — one per observation — so you can see how the partial derivative \(\partial \hat E[Y \mid X] / \partial X_j\) varies across the covariate space.
- Reports closed-form analytical standard errors for those marginal effects, which the package summarizes as pointwise CIs and “average marginal effects” (AMEs).
In short: a flexible regression that still tells you a coefficient.
The setup
KRLS represents the unknown regression function as a weighted sum of Gaussian kernels — one centered on each observation:
\[\hat f(x) \;=\; \sum_{i=1}^n c_i \,K(x, x_i) \qquad K(x, x_i) \;=\; \exp\!\left(-\frac{\|x - x_i\|^2}{\sigma^2}\right)\]The coefficients \(c\) minimize a Tikhonov-regularized least-squares objective,
\[\min_c \;\bigl\| Y - K c \bigr\|^2 + \lambda \, c^\top K c,\]where \(K\) is the \(n \times n\) kernel matrix and \(\lambda\) is a regularization parameter chosen by leave-one-out cross-validation. The closed-form solution is \(\hat c = (K + \lambda I)^{-1} Y\), and the implied \(\hat f\) lies in the reproducing kernel Hilbert space (RKHS) spanned by the Gaussian kernels.
The pointwise derivative \(\partial \hat f / \partial X_j\) at any data point comes out of \(\hat c\) in closed form. This is the headline output: not a single coefficient per predictor, but a distribution of marginal effects across observations.
Worked example
A simple simulated example showing what KRLS gives you that OLS doesn’t.
In R
library(KRLS)
set.seed(1)
n <- 200
x1 <- runif(n, -2, 2); x2 <- runif(n, -2, 2)
# True surface: nonlinear in x1, modulated by x2
y <- sin(x1) + 0.5 * x2 * (x1 > 0) + rnorm(n, sd = 0.3)
X <- cbind(x1, x2)
# Fit
fit <- krls(X = X, y = y)
# Pointwise marginal effects: one row per observation, one column per X
head(fit$derivatives)
# Average marginal effects (with analytical SEs)
summary(fit)
# Plot the distribution of marginal effects across observations
plot(fit)
For each predictor, summary(fit) reports:
- The average marginal effect (mean of pointwise derivatives).
- A standard error computed from the analytical vcov of \(\hat c\).
- Quartiles of the pointwise derivatives so you can see whether the effect is constant (boring) or varies across the covariate space (interesting).
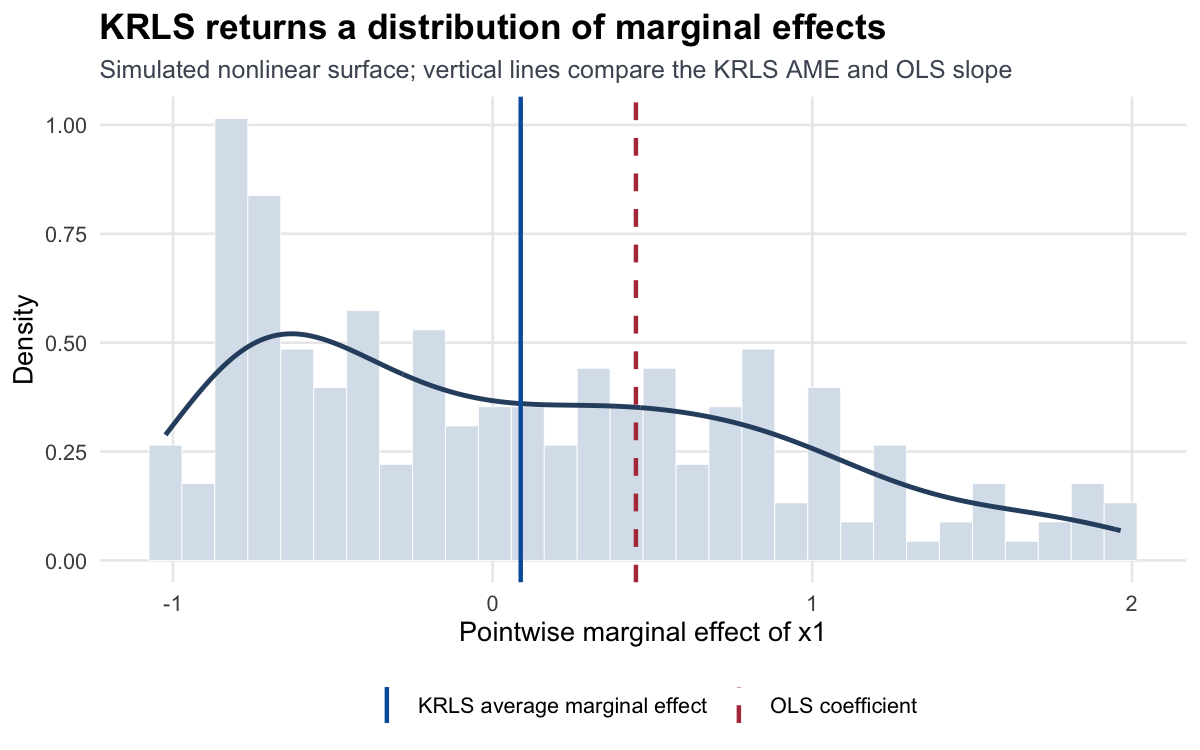
In this example the AME of x1 will be near zero — averaging \(\sin'(x_1) = \cos(x_1)\) over a symmetric range cancels — but the quartiles will show that the pointwise effect ranges from −1 to +1. That’s exactly the heterogeneity you’d miss with OLS.
In Stata
use mydata.dta, clear
krls y x1 x2
// Pointwise marginal effects are stored automatically; access via
matrix list e(derivatives)
// Plot heterogeneity in the marginal effect of x1
krls_plot x1
The Stata command shares the same algorithm and output structure as the R package; flags and defaults are documented in help krls.
Reading the output
Three things to look at:
1. Average marginal effects (AMEs). The analogue of an OLS coefficient. If the AME is large and the pointwise derivatives are all close to it, the relationship is roughly linear and KRLS gives you the same conclusion as OLS — but with a check that linearity held.
2. Heterogeneity in pointwise derivatives. This is the unique KRLS contribution. If pointwise derivatives are far from the AME for many observations, the marginal effect varies meaningfully across the covariate space. That’s substantive: the average effect masks heterogeneity. plot(fit) shows the distribution.
3. Pre/post-fit diagnostics.
fit$R2 # in-sample R²
fit$Looloss # leave-one-out CV loss at the chosen lambda
fit$lambda # regularization parameter (chosen via LOOCV)
fit$sigma # kernel bandwidth (default p; set sigma= for sensitivity)
If \(R^2\) is very high but the LOO loss is also high, you may be overfitting — try a larger \(\lambda\) via lambdasearch() or constrain the kernel bandwidth.
Minimum reporting checklist
When reporting a KRLS fit, include at least:
- average marginal effects with analytical standard errors;
- quartiles or a plot of the pointwise marginal effects;
- in-sample \(R^2\) and leave-one-out loss;
- the selected regularization parameter and kernel bandwidth;
- sensitivity checks for bandwidth or predictor scaling when results hinge on heterogeneity.
Choosing among alternatives
KRLS is in the family of nonparametric regression methods. Pick by what you need:
- Want pointwise marginal effects with closed-form SEs? KRLS is designed for this.
- Want fast prediction on big data? Random forests or gradient boosting (
ranger,xgboost) scale to millions of rows where KRLS does not. They give you SHAP values for interpretation, which approximate KRLS’s pointwise derivatives but with bootstrap-based uncertainty. - Have only a handful of nonlinear features in mind? GLM with splines (
mgcv) is a lighter tool that’s faster and usually sufficient. - Care most about the average effect, not heterogeneity? OLS with regression adjustment (and a check that linearity is reasonable) gets you there with a tiny fraction of the compute.
Common pitfalls
- n³ memory and runtime. KRLS solves an \(n \times n\) linear system. For \(n > \sim 5{,}000\), the cost becomes prohibitive on a single machine. The
bigKRLSpackage (now archived on CRAN) ported the heavy lifting to C++ via Rcpp/RcppArmadillo for a ~5–10× constant-factor speedup, but the asymptotic \(\mathcal O(n^3)\) scaling is unchanged. For genuinely large problems, consider Nyström approximation or random Fourier features (not currently inKRLSitself). - Bandwidth choice. The default is \(\sigma^2 = p\) where \(p\) is the number of predictors. This works well in many cases but is worth checking. Smaller \(\sigma\) makes the kernel sharper (higher variance, lower bias); larger \(\sigma\) smooths more. Refit with
sigma = ...and compare \(R^2\) + LOO loss. - Standardization matters. Gaussian kernels are scale-sensitive. Standardize predictors before fitting, or pass them as a matrix with comparable scales. The package does this internally by default.
- Marginal effects average over the sample, not the population. The AME of a covariate is the mean pointwise derivative across your data points. If your sample is selected, the AME reflects that selection.
- Categorical predictors. KRLS expects numeric predictors. For categories, expand to dummies before passing to
krls(). The package will return one marginal effect per dummy.
When not to use KRLS
- Very large \(n\). \(n > \sim 5{,}000\) on a typical machine becomes painful. Use a different method (or wait for the next generation of
KRLSwith Nyström approximation, planned). - You only care about prediction. If you don’t need pointwise marginal effects, gradient boosting will be faster and often more accurate.
- You need extrapolation outside the data. KRLS is non-parametric in the data range; outside it, predictions revert toward zero (in centered coordinates). Don’t ask it to extrapolate.
References
- Hainmueller, J., and Hazlett, C. (2014). Kernel regularized least squares: Reducing misspecification bias with a flexible and interpretable machine learning approach. Political Analysis, 22(2), 143–168.
- Ferwerda, J., Hainmueller, J., and Hazlett, C. (2017). Kernel-Based Regularized Least Squares in R (KRLS) and Stata. Journal of Statistical Software, 79(3), 1–26.
See also
- Kernel ML Methods project page — package landing page with the latest release notes.
- The companion Stata package:
krls(see https://github.com/j-hai/krls-stata). - For very large problems, the (archived)
bigKRLSpackage on the CRAN archive offers an Rcpp/RcppArmadillo port; modern alternatives include Nyström approximation in custom code or random Fourier features.